OmicsBox 3.4
OmicsBox (previously Blast2Go) is a leading bioinformatics software developed by BioBam Bioinformatics S.L. that offers end-to-end NGS data analysis of genomes, transcriptomes, and metagenomes. It is designed to be user-friendly, efficient, and with a powerful set of tools to extract biological insights from omics data. The software is structured in different modules, each with a specific set of tools and functions designed to perform different types of analysis, such as de-novo genome assemblies, genetic variation analysis, differential expression analysis, and taxonomic classifications of microbiome data, including the functional interpretation and rich visualisations of results. The functional analysis module, which includes the popular Blast2GO annotation methodology makes OmicsBox particularly suited for non-model organism research. OmicsBox works out of the box on any standard PC or laptop with Windows, Linux, or Mac.
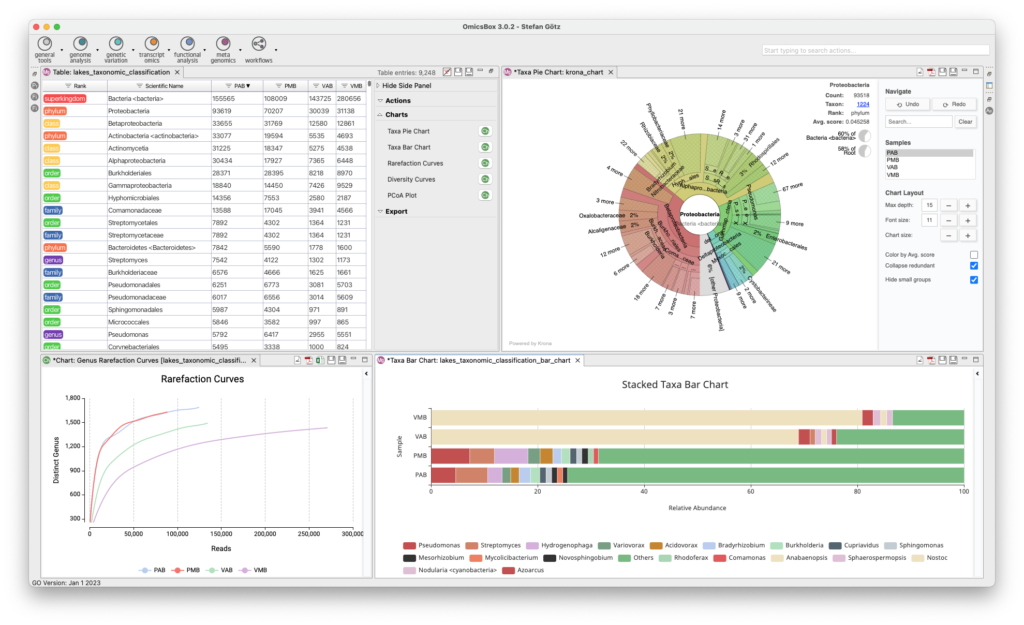
Functional Analysis
The Functional Analysis Module provides biological context as an analysis option. The Blast2GO methodology allows to flexibly assigns reliable functional functional labels to novel sequences through fast alignment, domain, and orthology information generated by cloud-based Blast, InterPro, and EggNog searches. Furthermore, enrichment approaches such as Fisher’s Exact Test and GSEA identify over- and underrepresented biological functions. The Combined Pathway Analysis identifies Reactome and KEGG pathways for any sequence set, allowing pathway enrichment calculation and providing rich visualizations for insights.
- Basic Sequence Analysis
- Blast2GO Functional Annotation
- Enrichment Analysis
- Pathway Analysis with KEGG
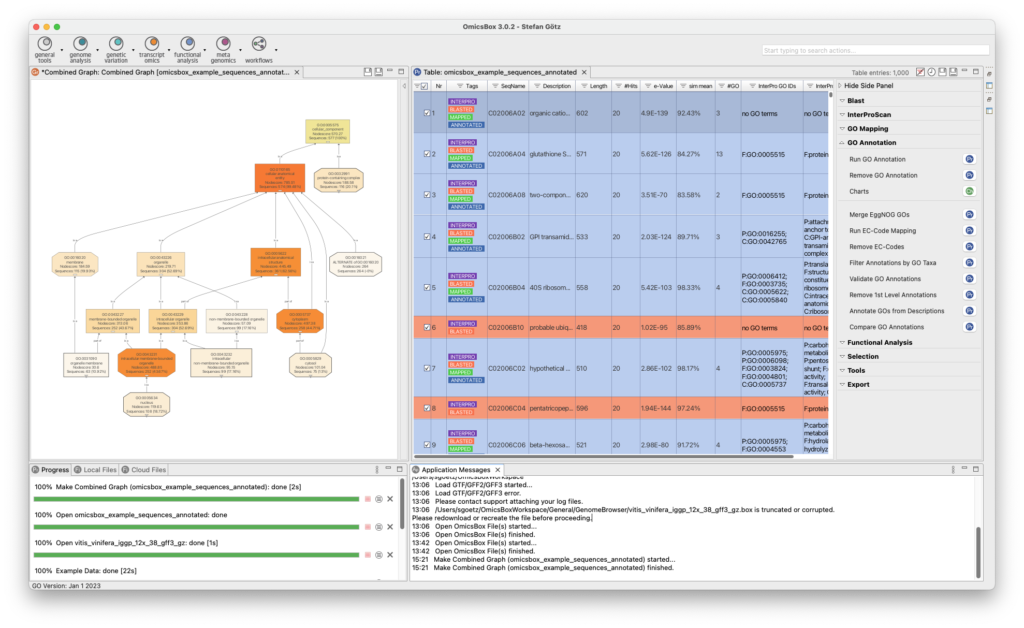
Genome Analysis
The Genome Analysis Module is an efficient and user-friendly toolset to characterizes and analyzes newly sequenced genomes. Quality control is performed using FastQC and Trimmomatic, followed by the assembly of whole genome sequences using popular algorithms such as ABySS, SPAdes, and Flye. BWA and Bowtie2 are used for read alignments, and tools for repeat identification and masking prior to gene prediction are also available. OmicsBox enables visualization of annotations through tracks that combine genome sequences, alignments, intron-exon structure, and variant data.
- Quality Control And Assessment
- Genome Mapping and Assembly
- Repeat Masking and Gene Finding
- Genome Curation Tools
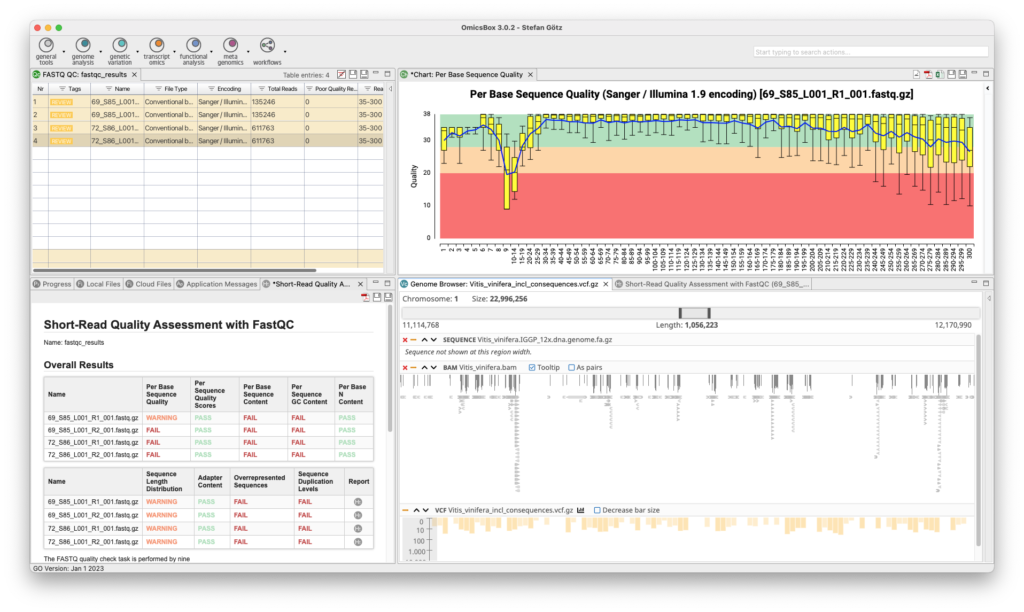
Genetic Variation
The Genetic Variation Module allows for variant calling, filtering, and annotation, as well as genome-wide association studies to associate genetic variations with a particular trait or disease. The module offers two analysis strategies for variant calling and filtering using BCFtools and FreeBayes. The resulting VCF file can be annotated via the Variant Effect Predictor from Ensembl, and guided genome-wide association studies can be executed to identify genetic variations associated with a particular trait. This tool combination outperforms alternatives in recent review studies.
- Fast Variant Calling and Filtering
- Supports GBS and WGS data
- Custom Variant Annotation
- Genome Wide Association Studies
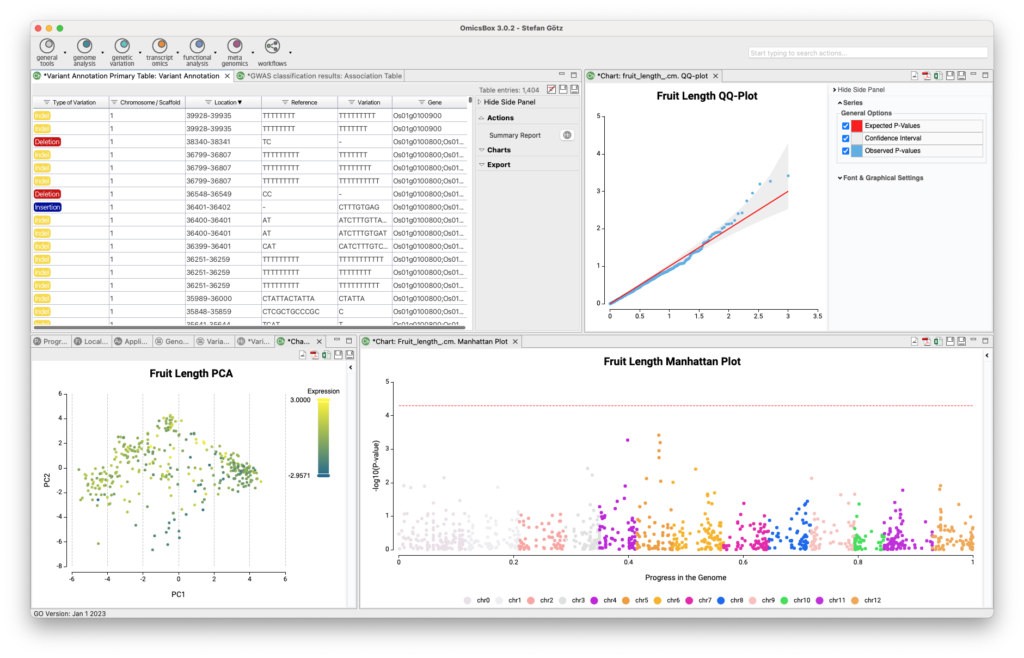
Transcriptomics
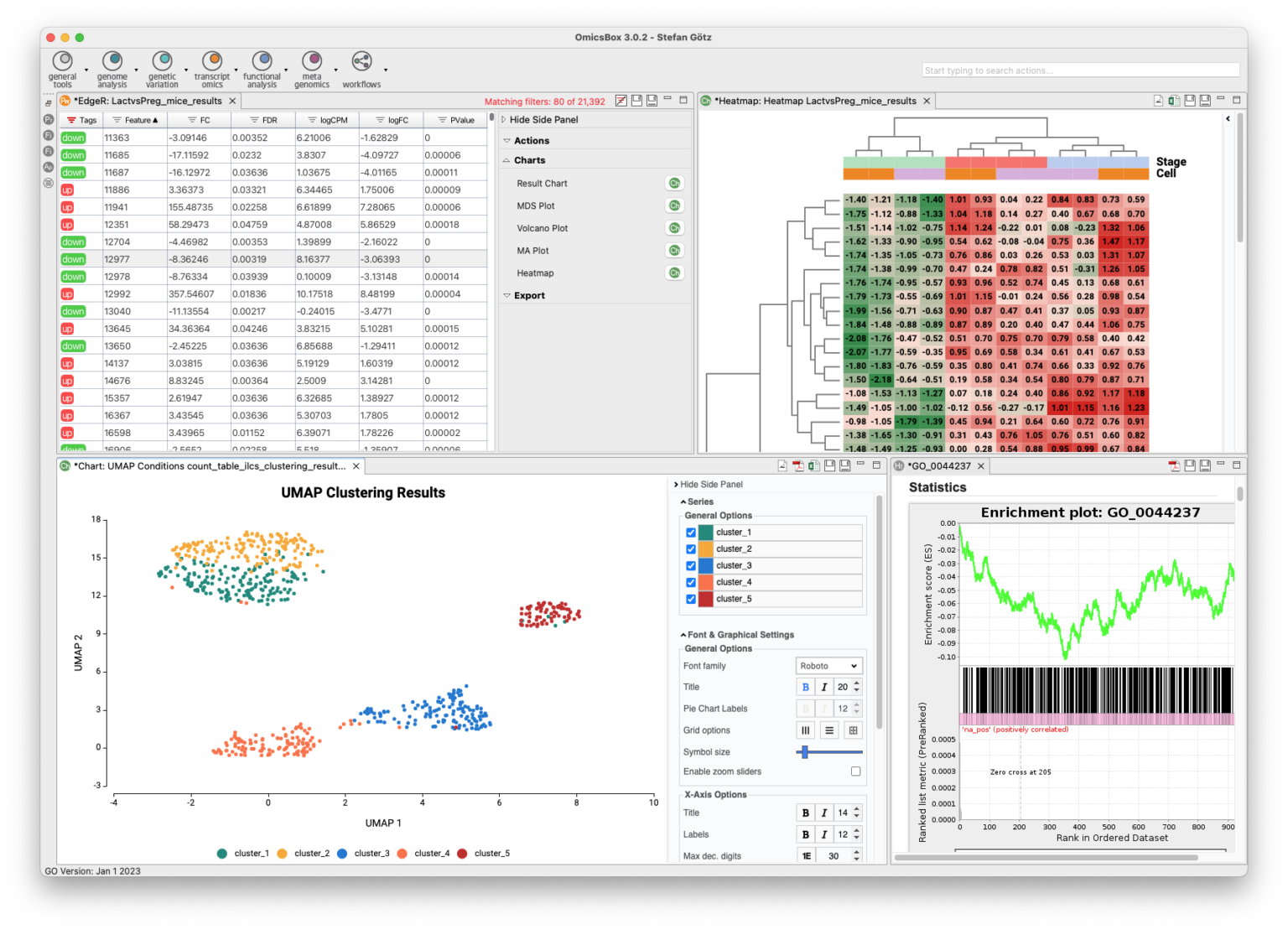
The Transcriptomics Module allows for processing RNA-Seq data from raw reads to functional analysis in a flexible and intuitive way. Quality control is followed by alignment to a reference genome using STAR or BWA or de-novo transcriptome assembly using Trinity. Coding regions can be predicted, and expression can be quantified using HTSeq or RSEM. Statistical packages like NOISeq, edgeR, or maSigPro detect differentially expressed genes. Single-Cell RNA-Seq tools identify cell groups and lineage trajectories. Enrichment analysis is integrated to identify over- and underrepresented biological functions easily.
- De-Novo Assembly and Alignment
- Differential Expression Analysis
- Long Read Transcriptomics
- Single-Cell RNA-Seq
CLICK HERE for more details and to request a TRIAL version.
Watch an Introductory Video about OmicsBox features.
Neogen Informatics Inc. is the sole authorized reseller of OmicsBox in Indian territory. Please request a quote if you would like to purchase the software.